This is old.
See the changelog for updates.
The XML input is outlined here. For those familiar with HTML language for web pages it should be easy to read. Accompanying the XML file should be the parratt.dtd file, which should not need to be altered. DTD files describe the rules for how the XML file should appear, to make sure that the number and order of records are valid.
The XML file begins with two header lines, that should always be the same. They specify the DTD file to look for. <?xml version="1.0"?>
<!DOCTYPE parratt SYSTEM "parratt.dtd">
The root level of XML tag is also always the same, and every option described below appears between the parratt tags. The parratt tag takes two optional attributes, type and do_fit, which must appear in quotes.
<parratt type="parratt" do_fit="no">
...
</parratt>
type attribute sets the type of simulation, whether normal parratt or polarized. The default value is parratt. When the do_fit attribute is set to no, the program only calculates the profile and reflectivity curves without doing any fitting. The default value is no.
The next section should is for parameters controlling the output. This elements in this section are optional. <output>
<reflectivity>reflectivity.txt</reflectivity>
<profile>profile.txt</profile>
<q_min> 0.000</q_min>
<q_max> 0.300</q_max>
<num_q>200</num_q>
<num_z>200</num_z>
</output>
num_q points from q_min to q_max. The profile curve is calculated at num_z points between limits determined by the program. Shown above are the default values.
The next section is for parameters controlling the fitting. This section is skipped if no fitting is to be done. <fit>
<data>reflectivity.dat</data>
<weight>WEIGHT_NONE</weight>
<type>LEAST_SQUARES</type>
<algo>LMSDR</algo>
</fit>
parratt mode. For polarized fitting of two data sets, use these tags instead: <data_up>data_up.dat</data_up>
<data_down>data_down.dat</data_down>
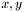 or three columns,
or three columns, 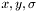 . The second tag describes what type of weights are be used for the fitting, in terms of the uncertainty in the
. The second tag describes what type of weights are be used for the fitting, in terms of the uncertainty in the 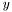 measurements,
measurements, 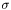 . The choices are
. The choices are WEIGHT_NONE | WEIGHT_R | WEIGHT_SQRT | WEIGHT_ERR
WEIGHT_NONE sets 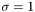 . Choosing
. Choosing WEIGHT_R or WEIGHT_SQRT sets 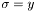 or
or 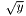 , respectively. The final choice
, respectively. The final choice WEIGHT_ERR is to use the reported error from the data file, which should be in the third column.
There are three different fitting method types, some with optional algorithms. Everything available in the Gnu Scientific Library version 1.6 was included.
Type: LEAST_SQUARES
Algo: LMSDR
Algo: LMDR
Type: MULTIDIM_MINIMIZAION
Algo: CONJUGATE_FR
Algo: CONJUGATE_PR
Algo: VECTOR_BFGS
Algo: STEEP_DESC
Type: SIMULATED_ANNEAL
Algo: SIMULATED_ANNEAL
The least squares algorithms are by far the most robust choice. I have found that although the multidimensional minimization use the computed first derivatives to pick the direction of optimization, the step size is not always as optimal. I have also used simulated annealing to gain insights into the variation of the parameters, through repeatedly running simulations from a already good fit as a starting point.
The last section is the model section. All data relating to the model falls between he model XML tags.
A general layer has five or seven quantities, depending on whether polarization is included.
<layer>
<name>SiO2</name>
<thik fix="0">11.000</thik>
<sigz fix="1"> 1.000</sigz>
<rsld fix="1"> 8.000</rsld>
<isld fix="1"> 0.000</isld>
<rmag fix="1"> 8.000</rmag>
<imag fix="1"> 0.000</imag>
</layer>
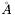 for the thickness (
for the thickness (thik) and roughness (sigz), and 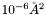 for the scattering length densities (SLD). The nuclear SLD is given by
for the scattering length densities (SLD). The nuclear SLD is given by rsld, and for the magnetic SLD as rmag. The imaginary part of either the SLD is optional. The order of these parameters is, I think, important, since it's dictated by the DTD.
The way things currently work, the program will read, and use, the rmag value in the calculation, regardless of the choice of parratt or polarization as the simulation type. The real purpose of the polarization type is just to fit two data sets at the same time. In that case, the value of rmag is used correctly for both spin up and down neutrons.
Therefore, if you do not want the magnetic SLD considered in your parratt calculation, do not include it in the layer. On the other hand, if you want to fit only one data set of polarized neutron data, you might include it. It will be used in the spin up case only, which is 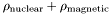 . Obviously, to use
. Obviously, to use rmag in the spin down case, make the value negative.
In order to hold a parameter fixed in the fit, set the fix attribute to 1, otherwise set to 0. This is ignored if not fitting. The fix attribute is optional, and the default is to hold the parameter fixed.
To make the layered strucutre easier to read in XML format, the bulk and substrate layers have their own tags, but follow the same parameters as any layer.
<bulk>
...
</bulk>
<substrate>
...
</substrate>
<substrate> or <bulk>. Remember to use the text "n/a" in place of numbers for <sigz> in the first layer, and for <thik> in the first and last layers, since these quantities don't have meaning in these layers.
Repeating groups of layers can be represented by enclosing the layers in a repeat tag.
<repeat num="3">
<layer>
...
</layer>
<layer>
...
</layer>
</repeat>
To simulate incomplete coverage of the substrate by the film, a coverage paramter can be set.
<coverg fix="1"> 1.000</coverg>
Lastly in the model section is the optional background and instrumental resolution.
<backgd fix="1"> 0.000</backgd>
<resoln type="dq"> 0.000</resoln>
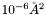 . The instrumental resolution requires the choice of whether
. The instrumental resolution requires the choice of whether 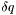 or
or 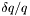 is constant by specifying the type as
is constant by specifying the type as dq or dq/q. See calcReflectivity() for more information. Without these tags, the default values assume no background or resolution smearing.
Included here is a complete XML file. <?xml version="1.0" encoding="UTF-8"?>
<!DOCTYPE parratt SYSTEM "parratt.dtd">
<parratt type="parratt" do_fit="no">
<output>
<reflectivity>reflectivity.txt</reflectivity>
<profile>profile.txt</profile>
<q_min> 0.000</q_min>
<q_max> 0.080</q_max>
<num_q>200</num_q>
<num_z>200</num_z>
</output>
<fit>
<data>reflectivity.dat</data>
<!--
<data_up>reflup.dat</data_up>
<data_down>refldn.dat</data_down>
-->
<weight>WEIGHT_R</weight>
<type>LEAST_SQUARES</type>
<algo>LMSDR</algo>
</fit>
<model>
<bulk>
<name>solvent bulk</name>
<thik fix="1">n/a</thik>
<sigz fix="1">n/a</sigz>
<rsld fix="1"> 0.000</rsld>
</bulk>
<layer>
<name>film</name>
<thik fix="0">460.00</thik>
<sigz fix="0"> 18.51</sigz>
<rsld fix="0"> 2.348</rsld>
</layer>
<layer>
<name>oxide</name>
<thik fix="1">18.000</thik>
<sigz fix="1"> 8.000</sigz>
<rsld fix="1"> 3.475</rsld>
<isld fix="1"> 1.049E-5</isld>
</layer>
<substrate>
<name>silicon</name>
<thik fix="1">n/a</thik>
<sigz fix="1"> 4.000</sigz>
<rsld fix="1"> 2.073</rsld>
<isld fix="1"> 2.376E-5</isld>
</substrate>
<coverg fix="1"> 1.000</coverg>
<backgd fix="1"> 0.000</backgd>
<resoln type="array-dq/q">resolution.txt</resoln>
</model>
</parratt>
<!ELEMENT parratt (output?,fit?,model)>
<!ATTLIST parratt type CDATA "parratt">
<!ATTLIST parratt do_fit CDATA "no">
<!-- output section -->
<!ELEMENT output (reflectivity?,profile?,q_min?,q_max?,num_q?,num_z?)>
<!ELEMENT reflectivity (#PCDATA)>
<!ELEMENT profile (#PCDATA)>
<!ELEMENT q_min (#PCDATA)>
<!ELEMENT q_max (#PCDATA)>
<!ELEMENT num_q (#PCDATA)>
<!ELEMENT num_z (#PCDATA)>
<!-- fit section -->
<!ELEMENT fit (data?,data_up?,data_down?,weight?,(type|algo)*)>
<!ELEMENT data_up (#PCDATA)>
<!ELEMENT data_down (#PCDATA)>
<!ELEMENT data (#PCDATA)>
<!ELEMENT weight (#PCDATA)>
<!ELEMENT type (#PCDATA)>
<!ELEMENT algo (#PCDATA)>
<!-- model section -->
<!ELEMENT model ((bulk|substrate),(repeat|layer)*,(bulk|substrate),coverg?,backgd?,resoln?)>
<!ELEMENT bulk (name,thik?,sigz?,rsld,isld?,rmag?,imag?)>
<!ELEMENT layer (name,thik,sigz,rsld,isld?,rmag?,imag?)>
<!ELEMENT repeat (layer+)>
<!ATTLIST repeat num CDATA "1">
<!ELEMENT substrate (name,thik?,sigz?,rsld,isld?,rmag?,imag?)>
<!ELEMENT name (#PCDATA)>
<!ELEMENT thik (#PCDATA)>
<!ATTLIST thik fix CDATA "1">
<!ELEMENT sigz (#PCDATA)>
<!ATTLIST sigz fix CDATA "1">
<!ELEMENT rsld (#PCDATA)>
<!ATTLIST rsld fix CDATA "1">
<!ELEMENT isld (#PCDATA)>
<!ATTLIST isld fix CDATA "1">
<!ELEMENT rmag (#PCDATA)>
<!ATTLIST rmag fix CDATA "1">
<!ELEMENT imag (#PCDATA)>
<!ATTLIST imag fix CDATA "1">
<!ELEMENT coverg (#PCDATA)>
<!ATTLIST coverg fix CDATA "1">
<!ELEMENT backgd (#PCDATA)>
<!ATTLIST backgd fix CDATA "1">
<!ELEMENT resoln (#PCDATA)>
<!ATTLIST resoln type CDATA "dq/q">
Generated on Wed Mar 14 13:24:56 2007 for Parratt by
 1.4.7
1.4.7
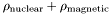 . Obviously, to use
. Obviously, to use 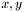 or three columns,
or three columns, 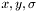 . The second tag describes what type of weights are be used for the fitting, in terms of the uncertainty in the
. The second tag describes what type of weights are be used for the fitting, in terms of the uncertainty in the 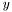 measurements,
measurements, 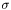 . The choices are
. The choices are 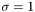 . Choosing
. Choosing 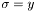 or
or 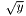 , respectively. The final choice
, respectively. The final choice 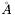 for the thickness (
for the thickness (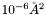 for the scattering length densities (SLD). The nuclear SLD is given by
for the scattering length densities (SLD). The nuclear SLD is given by 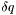 or
or 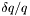 is constant by specifying the type as
is constant by specifying the type as  1.4.7
1.4.7